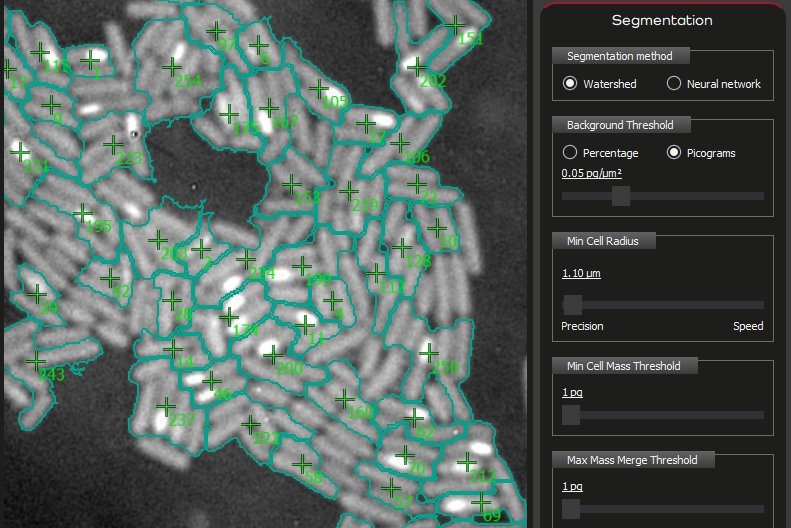
Telight introduces AI-assisted cell segmentation within the Q-Phase platform, extending the SophiQ software with a model-based approach to one of the most critical steps in quantitative live-cell analysis.
Cell segmentation has traditionally relied on threshold-based methods requiring manual parameterization. While widely adopted, these approaches introduce variability between users and sensitivity to dataset conditions. In practice, segmentation outcomes depend on parameter selection, background stability, and experimental duration, which make reproducibility difficult to maintain – particularly in long-term time-lapse experiments. The new functionality replaces threshold-based segmentation with neural network models trained on Q-Phase data. This enables consistent segmentation across entire datasets without the need for per-experiment parameter tuning, while keeping segmentation as an integrated part of the analytical workflow.
From a methodological perspective, this represents a transition from user-dependent parameterization to a model-driven approach. Because the segmentation is applied uniformly across all frames and positions, results become less sensitive to background fluctuations and more stable over time. This directly addresses segmentation drift, a common limitation in long-term experiments where traditional methods gradually lose consistency.
In addition to improving reproducibility, the model-based approach expands the range of analysable objects. Smaller cells, bacteria, and more complex morphologies can be detected more reliably, as segmentation is no longer constrained by fixed thresholds but instead based on learned cell boundaries.
An important aspect of this implementation is its integration within the Q-Phase system. Unlike external AI pipelines, where segmentation is performed as a separate post-processing step, Q-Phase incorporates AI-assisted segmentation directly into the SophiQ environment. This ensures that improvements in segmentation are immediately reflected in the quantitative outputs and eliminates the need for additional tools or data transfer between systems.
From a practical standpoint, the impact is immediate. Users spend less time on parameter tuning and manual correction, while achieving more consistent results across experiments and users. This improves both workflow efficiency and confidence in the data. The introduction of AI-assisted segmentation also establishes a foundation for further analytical capabilities. With segmentation stabilized, it becomes possible to build higher-level analysis on top of it, including detection of biological processes such as mitosis, cell death, and classification of cell populations.
Segmentation is not only a technical step. It is the point at which imaging data becomes measurable biology. By replacing subjective parameterization with a consistent, model-based approach, Q-Phase reduces a key source of variability and strengthens the reliability of quantitative live-cell experiments.
See how AI-assisted segmentation performs on your data and request demo now.